What Are Convolutional Neural Networks?
GENE 46100 — Unit 00
2026-03-24
What is a CNN?
A convolutional neural network (CNN) is a network architecture for deep learning that learns directly from data (images, sequences, …).
A CNN is made up of several layers that process and transform an input to produce an output.
Applications
- Scene classification
- Object detection & segmentation
- Image processing
- DNA sequence analysis (our focus!)
Three Key Concepts
Source: MathWorks — What Are CNNs?
- Local receptive fields
- Shared weights and biases
- Activation and pooling
1 — Local Receptive Fields
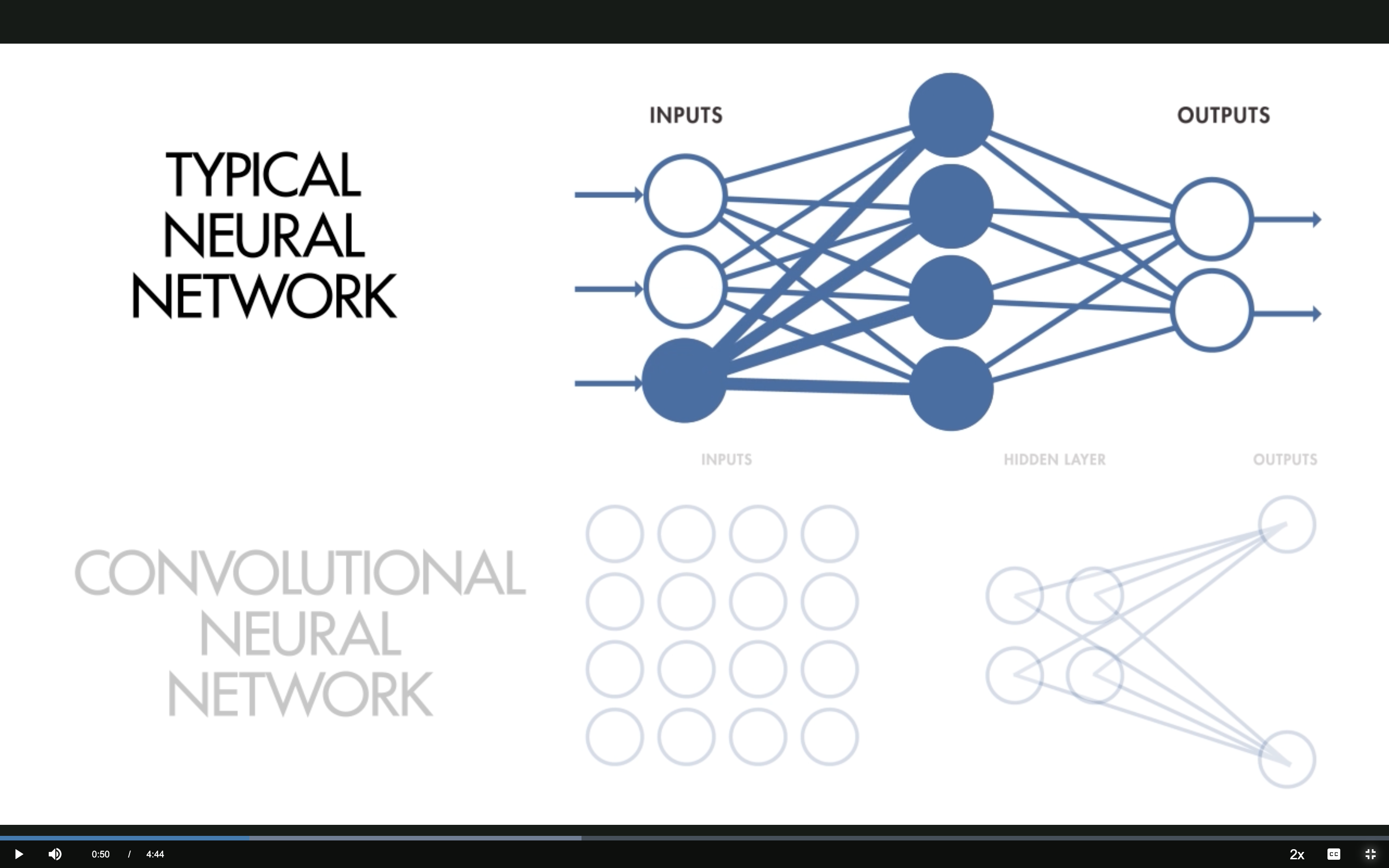
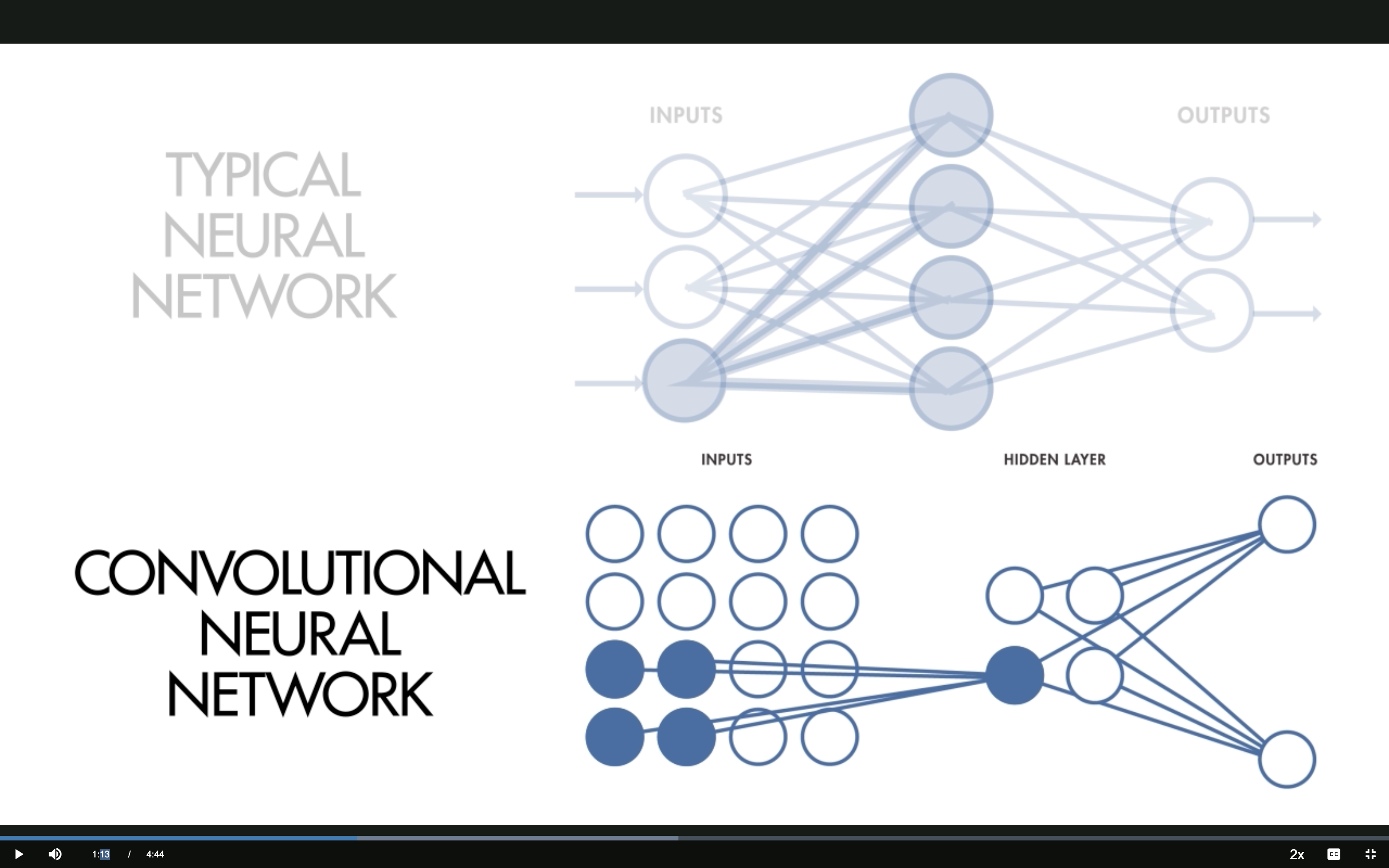
Source: MathWorks — What Are CNNs?
Typical NN: Fully Connected
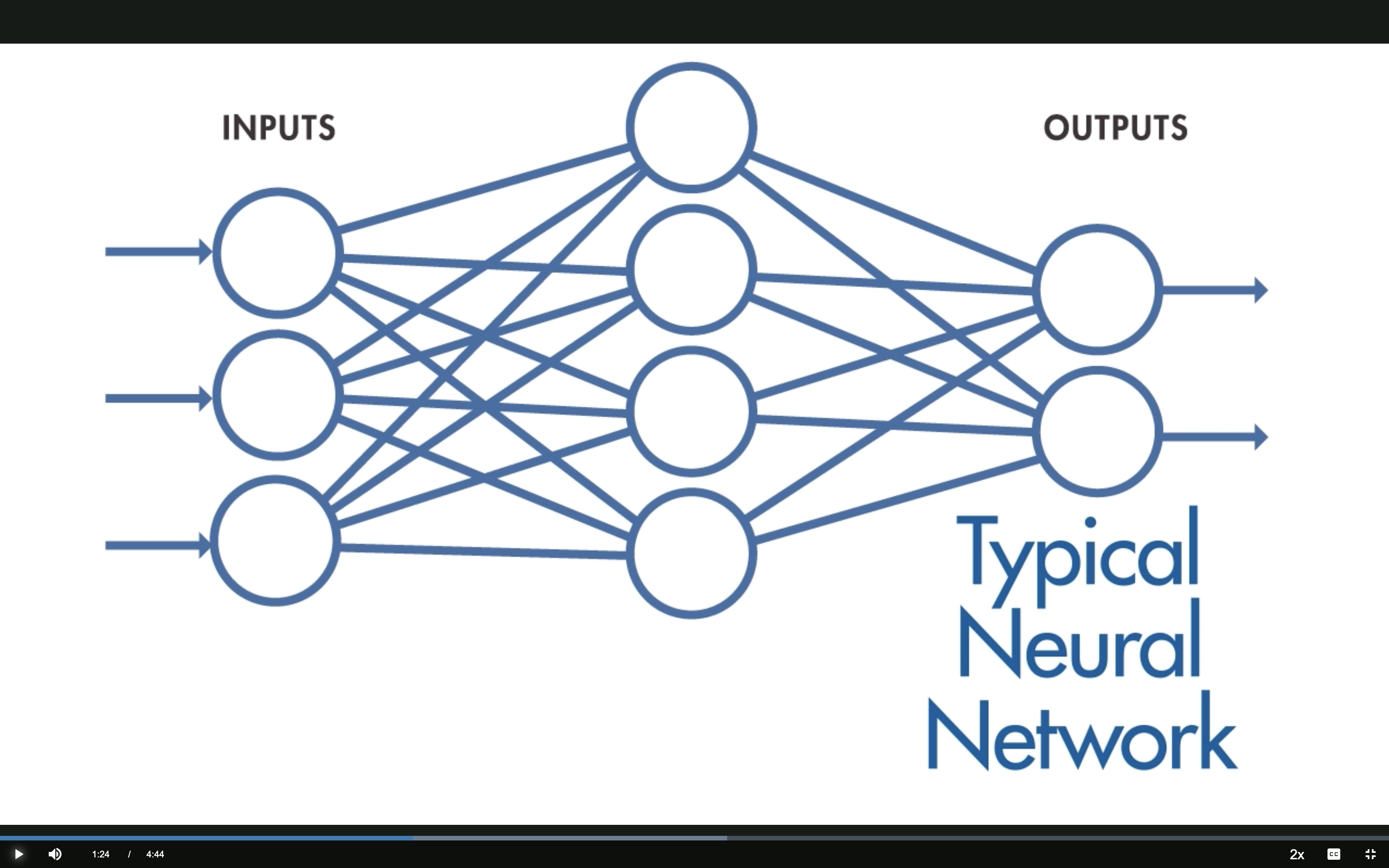
Every input neuron connects to every hidden neuron — each hidden neuron has its own set of weights.
Source: MathWorks — What Are CNNs?
Typical NN: Weights
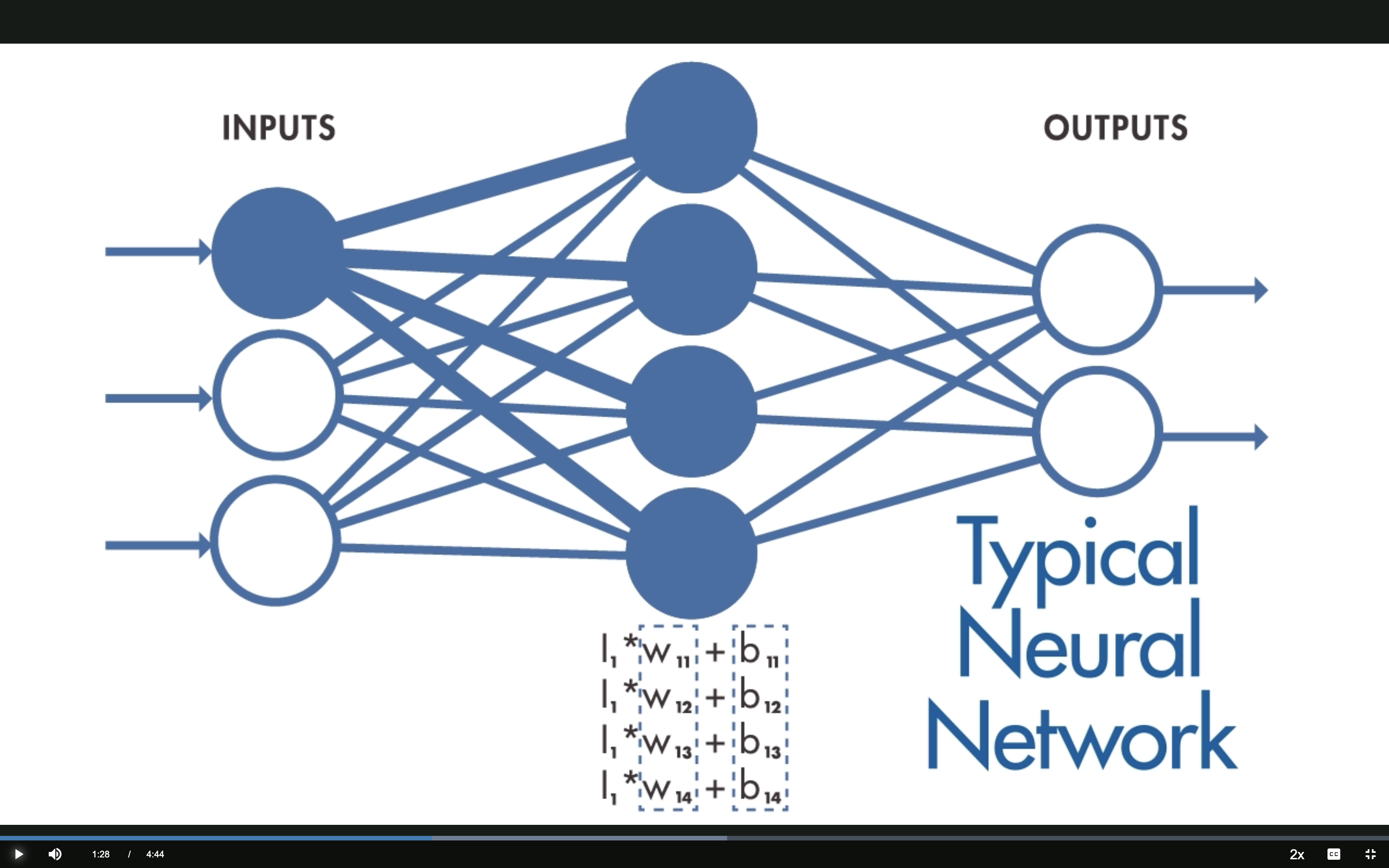
Hidden neuron output: \(h_j = \sum_i I_i \cdot W_{ij} + b_j\)
All \(N \times M\) weights are independent — expensive and no spatial structure.
Source: MathWorks — What Are CNNs?
CNN: Local Receptive Fields
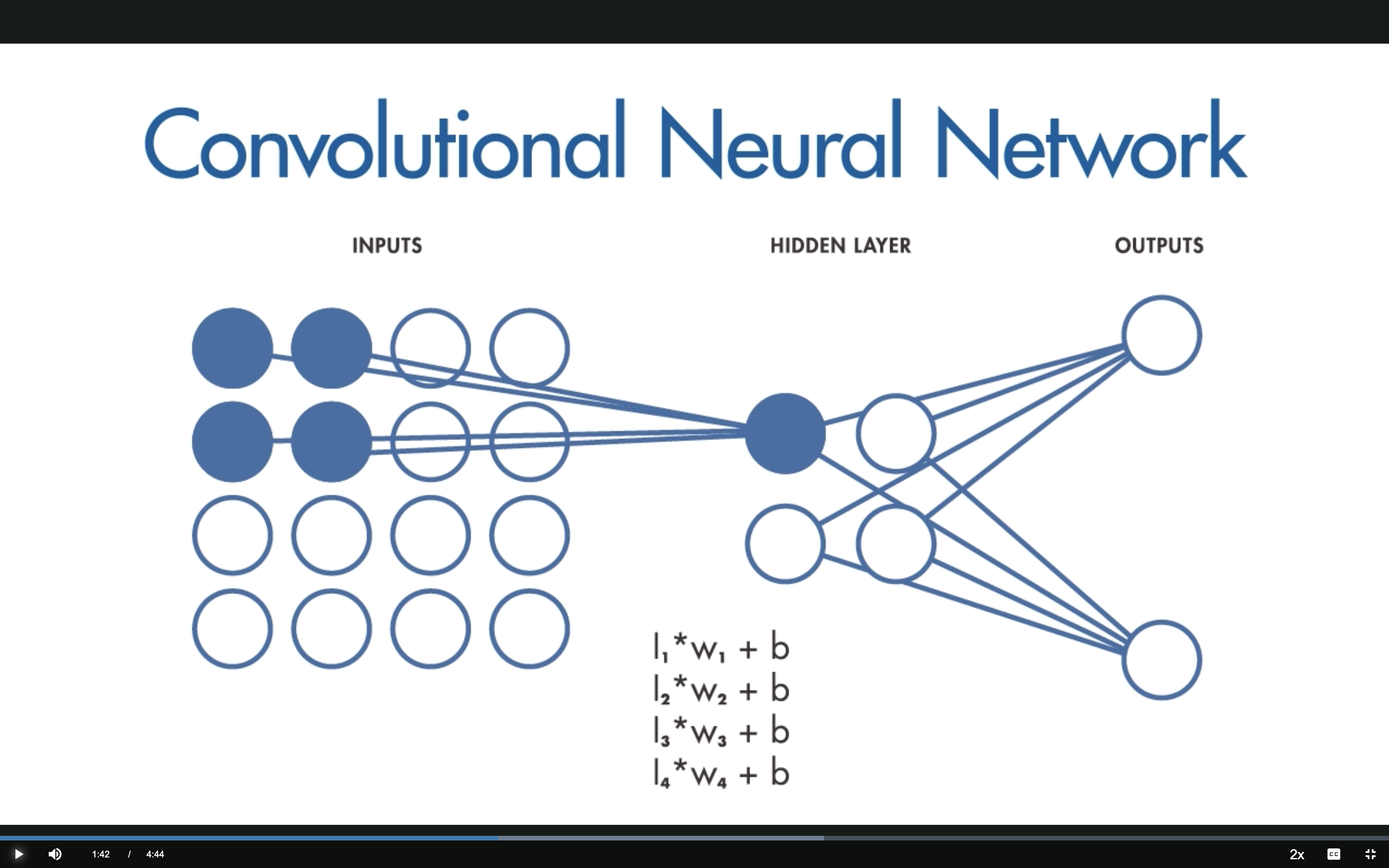
Only a small region of the input connects to each hidden neuron.
The hidden neuron computes: \(h = \sum_k I_k \cdot w_k + b\) using a small filter.
Source: MathWorks — What Are CNNs?
CNN: The Filter Slides
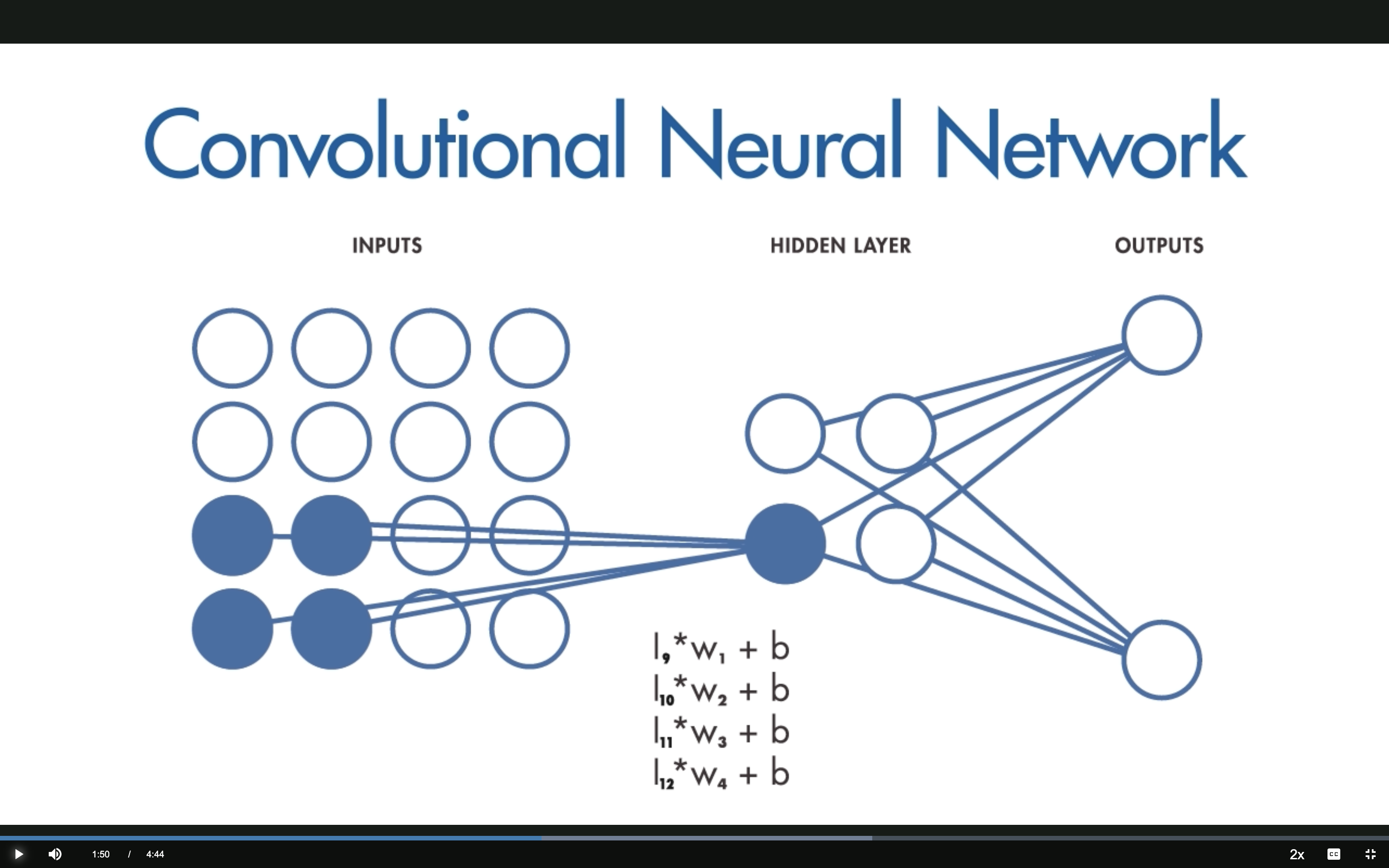
The same filter (same weights) slides across the input to create a feature map.
This sliding operation is convolution — hence the name convolutional neural network.
Source: MathWorks — What Are CNNs?
2 — Shared Weights and Biases
Key difference from a typical NN: the weights and biases are the same for all hidden neurons in a given layer.
This means every neuron in a layer is detecting the same feature (e.g., an edge or a blob) at different locations.
Compare the equations:
| Typical NN | CNN | |
|---|---|---|
| Weights | \(I_i \cdot W_{ij} + b_j\) (unique per neuron) | \(I_k \cdot w_k + b\) (shared) |
| Parameters | \(N \times M\) | Filter size only |
Source: MathWorks — What Are CNNs?
Translation Invariance
Because weights are shared, CNNs are tolerant to translation of objects in the input.
A network trained to recognize cats will find the cat wherever it appears in the image.
Genomics analogy: A CNN trained to recognize a TF binding motif will find it regardless of position in the DNA sequence.
Source: MathWorks — What Are CNNs?
3 — Activation and Pooling
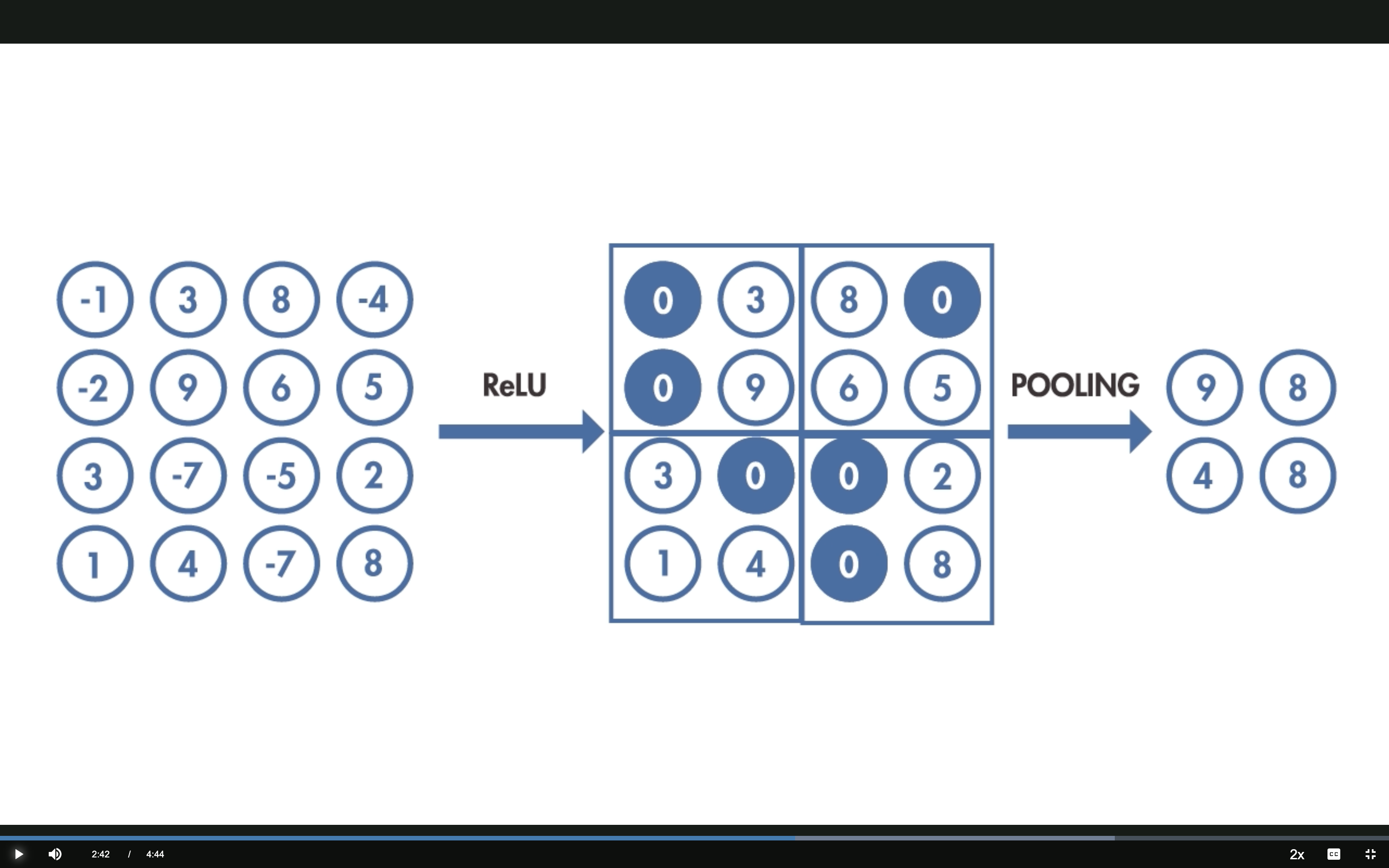
ReLU: \(f(x) = \max(0, x)\) — negative values become 0, positive values pass through.
Max Pooling: take the maximum value in each 2×2 block — reduces dimensionality while keeping the strongest activations.
Source: MathWorks — What Are CNNs?
Putting It All Together
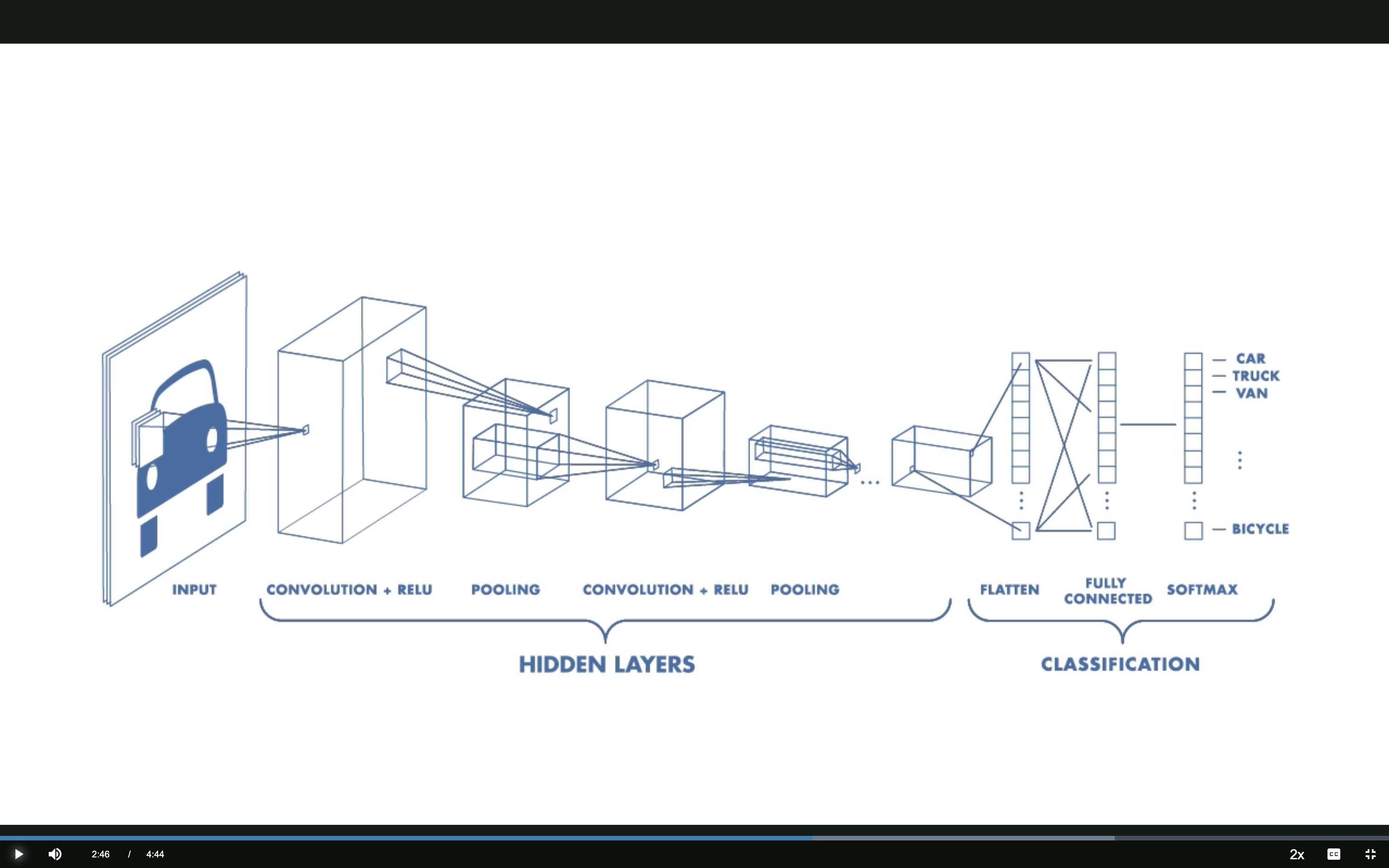
INPUT → CONV+RELU → POOLING → CONV+RELU → POOLING → FLATTEN → FULLY CONNECTED → SOFTMAX
Source: MathWorks — What Are CNNs?
Hierarchical Feature Learning
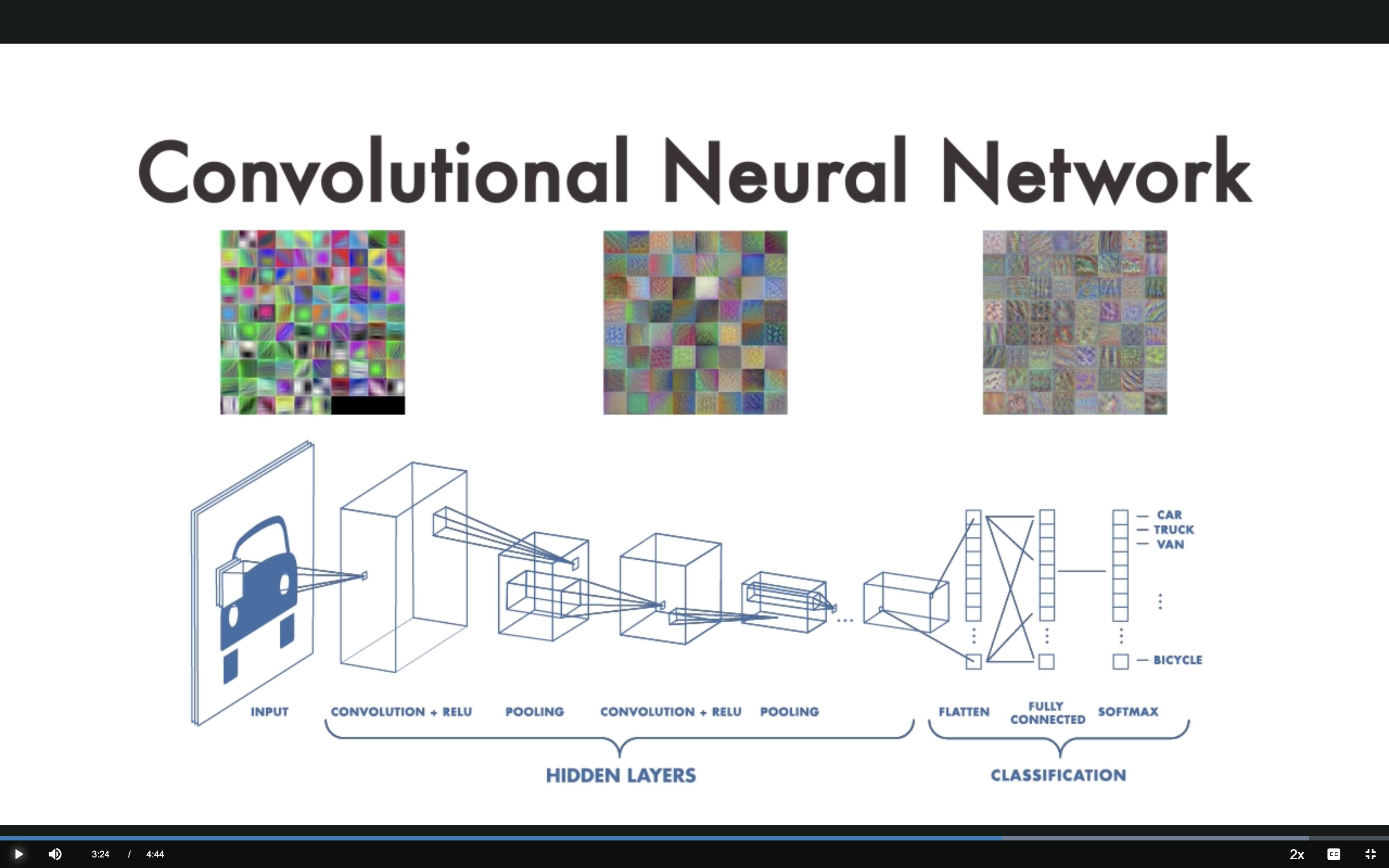
Each layer learns progressively more complex features:
| Layer depth | Vision | Genomics |
|---|---|---|
| Early layers | Edges, colors | Short k-mers |
| Middle layers | Textures, shapes | Motifs (e.g., TATA box) |
| Deep layers | Object parts, objects | Motif combinations, regulatory logic |
Source: MathWorks — What Are CNNs?
Three Ways to Use CNNs
Method 1 — Train from Scratch
Build and train the entire network on your own data.
Pros: Highly accurate, fully customized
Cons:
- Requires hundreds of thousands of labeled examples
- Significant computational resources (GPUs)
This is what we do in the TF binding prediction homework.
Source: MathWorks — What Are CNNs?
Method 2 — Transfer Learning
Use a model pre-trained on one task as the starting point for a related task.
Example: A CNN trained to recognize animals can be fine-tuned to distinguish cars from trucks.
Pros: Requires less data and compute
Genomics example: Fine-tune a model trained on one TF’s binding data to predict binding for a related TF.
Source: MathWorks — What Are CNNs?
Method 3 — Feature Extraction
Use a pre-trained CNN’s hidden layers to extract features, then train a simpler model on top.
A layer that learned to detect edges is broadly useful across many domains.
Pros: Requires the least data and compute
Genomics example: Use Enformer embeddings as features for downstream prediction tasks.
Source: MathWorks — What Are CNNs?
Summary
| Concept | Key Idea |
|---|---|
| Local receptive fields | Small sliding window over input |
| Shared weights | Same filter at every position → translation invariance |
| Activation (ReLU) | Introduce non-linearity |
| Pooling | Reduce dimensionality |
| Hierarchical features | Simple → complex across layers |
Summary
Three training strategies
- From scratch — most data, most flexibility
- Transfer learning — fine-tune a pretrained model
- Feature extraction — least data needed
Next Steps
- Hands-on: Build a CNN for TF binding prediction
- Key question: Can a CNN learn to detect regulatory motifs in DNA?
- We’ll use PyTorch to implement and train our CNN
GENE 46100 · Deep Learning in Genomics